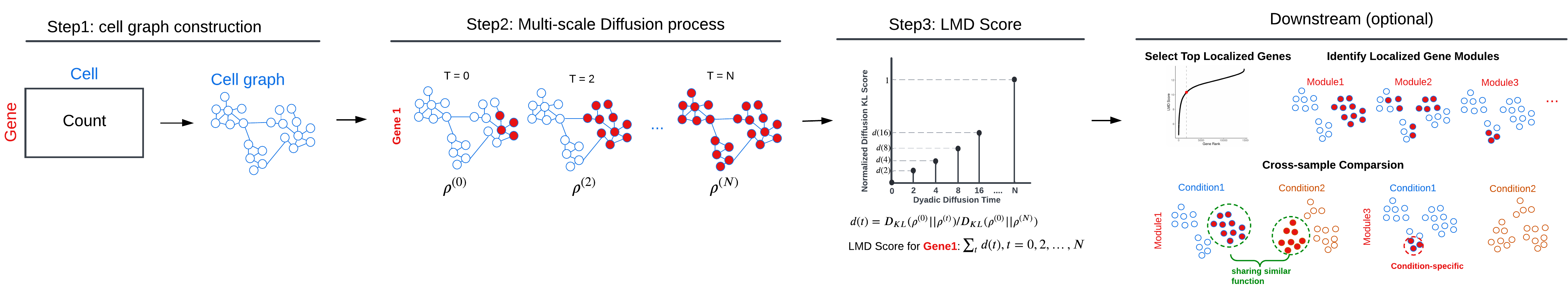
Localized Marker Detector (LMD) is a computational framework designed for the identification of gene expression markers localized to specific cell populations within single-cell RNA sequencing data. The major workflow of LMD comprises the following three main steps:
- Step1. Constructing a cell-cell affinity graph
- Step2. Diffusing the gene expression value across the cell graph
- Step3. Assigning a score to each gene based on the dynamics of its diffusion process
- Optional Downstream tasks
- Identifying gene modules and characterizing functional cell groups
- Cross-sample comparison
Installation
LMD can be installed in R as follows:
install.packages("devtools")
devtools::install_github("KlugerLab/LocalizedMarkerDetector")
library("LocalizedMarkerDetector")Docker Installation
Alternatively, we provide a Docker environment with LMD and all its dependencies pre-installed. The pre-built Docker image can be downloaded using:
and to run the docker container:
Here, change /data:/data to <your_local_data_directory>:/data. A rstudio will be on port 8029.
Example tutorial
Please check LMD tutorial.
References
References of LMD functions can be found here.
For additional information, please reference our paper:
Cluster-independent multiscale marker identification in single-cell RNA-seq data using localized marker detector (LMD) Ruiqi Li, Rihao Qu, Fabio Parisi, Francesco Strino, Hainan Lam, Jay S Stanley III, Xiuyuan Cheng, Peggy Myung, Yuval Kluger Communications Biology. 2025 Jul 16;8(1):1058. doi: https://doi.org/10.1038/s42003-025-08485-y